Notes
Sort by:| Metadata | Start Date | End Date | Comment |
|---|---|---|---|
|
CE02SHSM-RID26-08-SPKIRB000 |
There is no pressure variable in the .nc files. This should be a coordinate. By Lori Garzio, on 7/11/19 |
||
|
CE02SHSM-RID27-01-OPTAAD000 Deployment: 6 |
56% of the optical_absorption values for this deployment are nans, fill values, outside of global ranges, or outliers (+/- 5 SD). By Lori Garzio, on 6/28/19 |
||
|
CE02SHSM-RID27-01-OPTAAD000 |
There is no pressure variable in the .nc files. This should be a coordinate. By Lori Garzio, on 6/28/19 |
||
|
CE02SHSM-RID27-01-OPTAAD000 |
According to Roesler and Barnard (Optical proxy for phytoplankton biomass in the absence of photophysiology: Rethinking the absorption line height, Methods in Oceanography, 2013), “absorption meters are highly prone to biofouling, particularly biofilms which not only attenuate the collimated beam but also impact the scattering properties of the optical surfaces and tubes. In productive coastal waters biofouling can have significant impacts (i.e. 10% of the signal) within one to two weeks”. By Lori Garzio, on 6/28/19 |
||
|
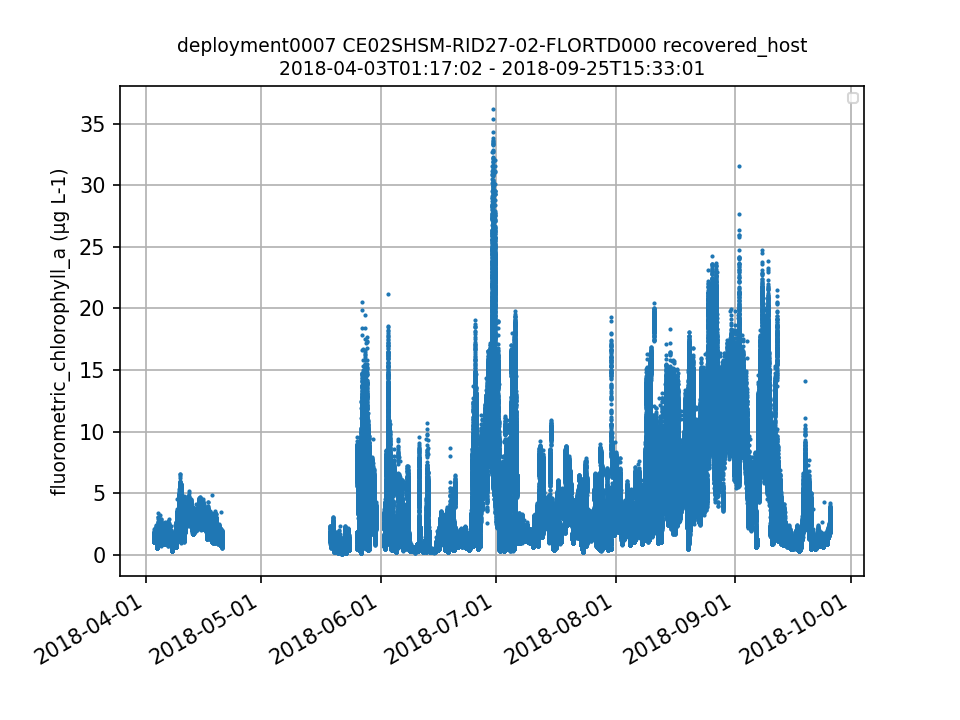
CE02SHSM-RID27-02-FLORTD000 Deployment: 7 |
|||
|
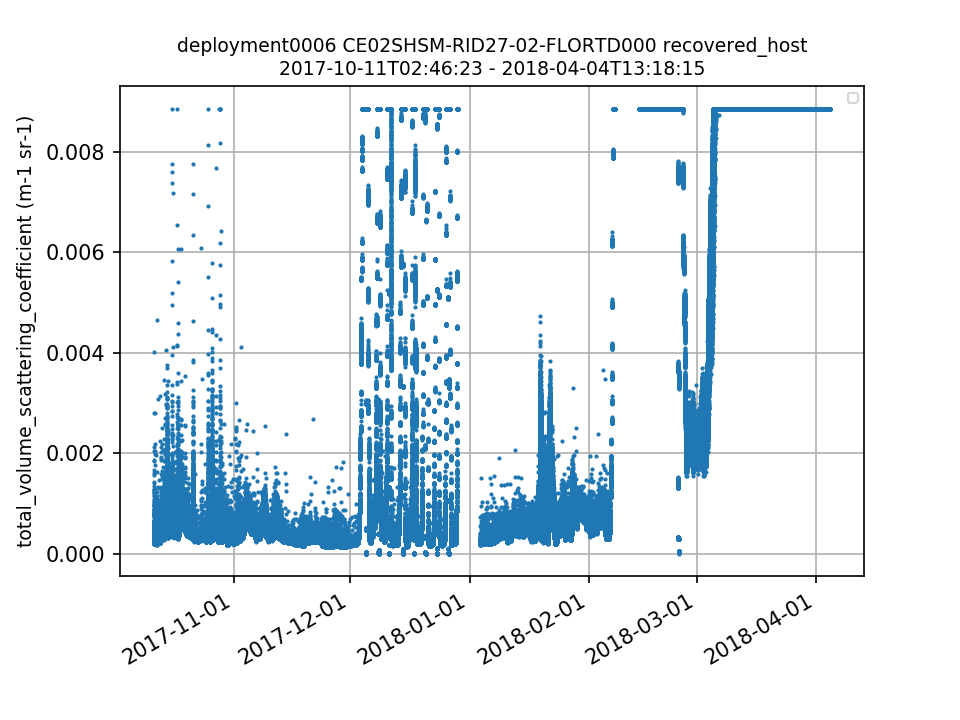
CE02SHSM-RID27-02-FLORTD000 Deployment: 6 |
12/1/17, 12:00 AM | 4/4/18, 1:57 PM |
total_volume_scattering_coefficient data show a repetitive pattern of increasing to the upper detection limit of the instrument and dropping to zero. A similar issue has been seen on CE06ISSM-RID16-02-FLORTD000 (see annotation ID 97). chl-a and CDOM data are also suspect. This dataset should be annotated. By Lori Garzio, on 7/3/19 |
|
CE02SHSM-RID27-02-FLORTD000 Deployment: 2 |
3/1/16, 12:00 AM | 5/16/16, 9:09 PM | |
|
CE02SHSM-RID27-02-FLORTD000 |
The variable pressure_depth (dbar) is an array of all fill values for every deployment. This should contain valid pressure data. By Lori Garzio, on 7/3/19 |
||
|
CE02SHSM-RID27-02-FLORTD000 Deployment: 5 |
The missing data for deployment 5 should be annotated. By Lori Garzio, on 3/26/19 |
||
|
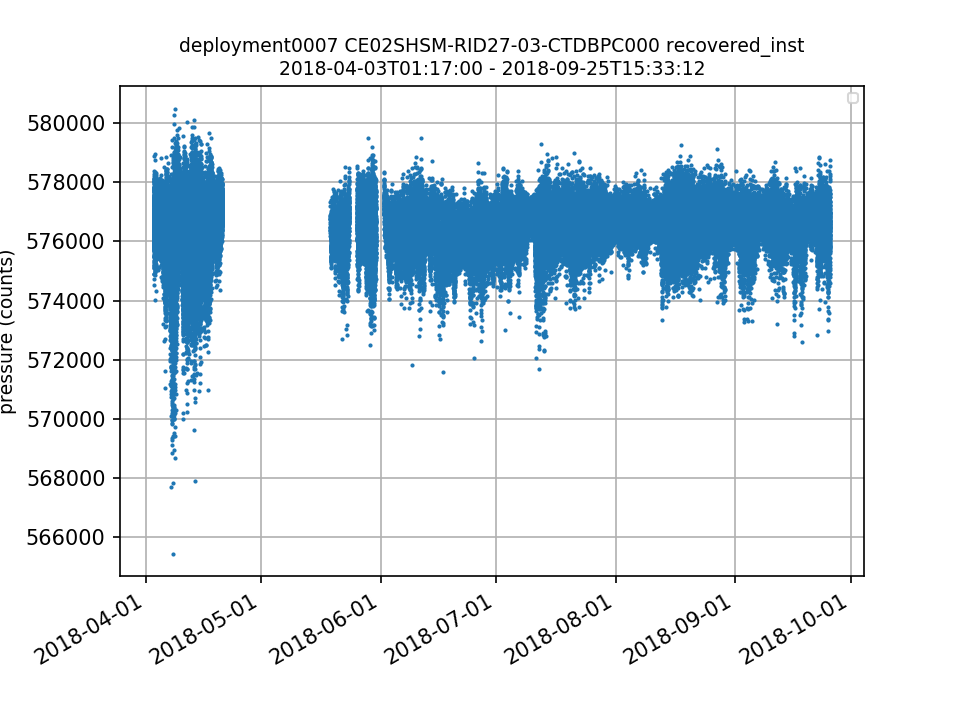
CE02SHSM-RID27-03-CTDBPC000 Deployment: 7 |
|||
|
CE02SHSM-RID27-03-CTDBPC000 Deployment: 6 |
10/10/17, 2:21 AM | 4/4/18, 1:57 PM |
Deployment 6 should be annotated to indicate why recovered_inst data aren't available. By Lori Garzio, on 3/26/19 |
|
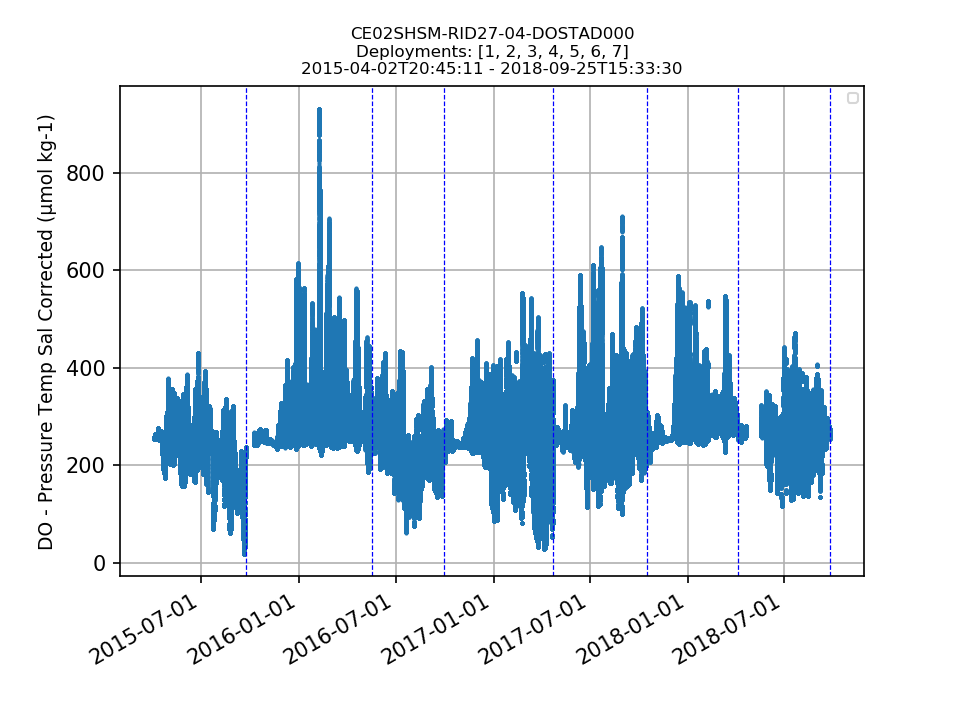
CE02SHSM-RID27-04-DOSTAD000 |
A substantial percentage of data for all deployments is outside of global ranges. It looks like the instruments biofouled shortly after each deployment (1-2 months) - the suspect data should be annotated for all deployments, and further analysis should be done using shipboard bottle data (which is outside the scope of this project). By Lori Garzio, on 3/26/19 |
||
|
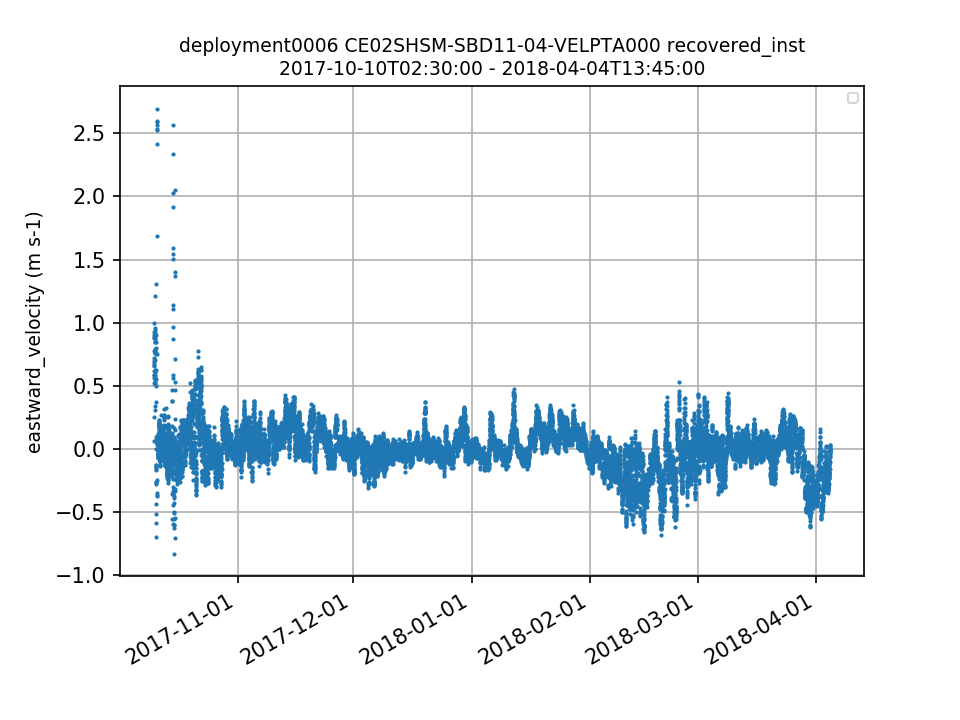
CE02SHSM-SBD11-04-VELPTA000 Deployment: 6 |
10/10/17, 2:21 AM | 10/20/17, 12:00 AM | |
|
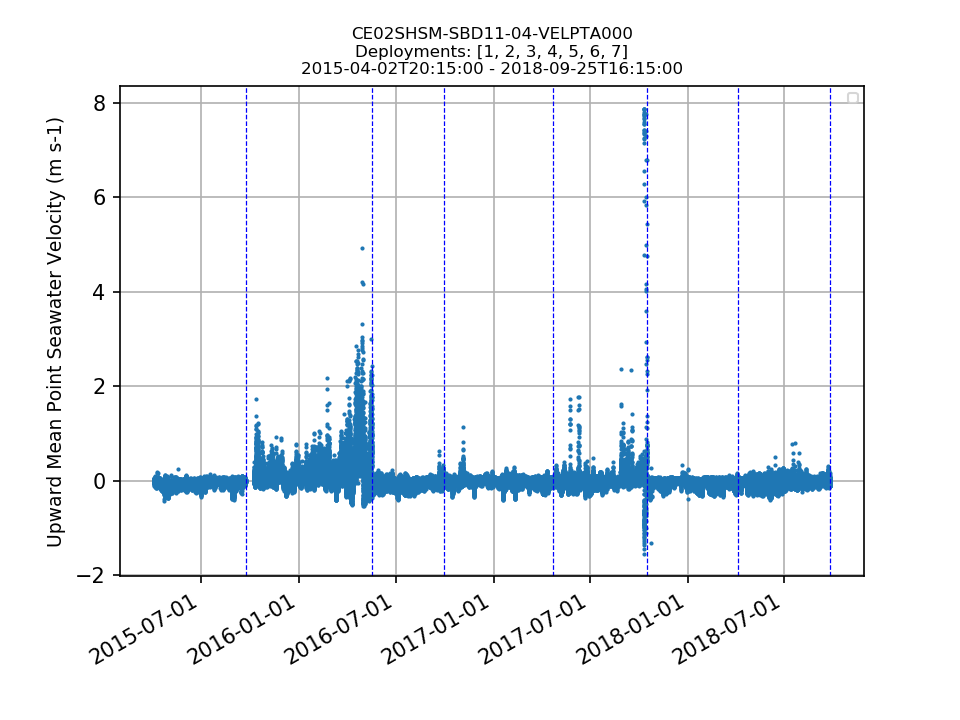
CE02SHSM-SBD11-04-VELPTA000 Deployment: 2 |
10/7/15, 9:10 PM | 5/16/16, 9:09 PM | |
|
CE02SHSM-SBD11-06-METBKA000 Deployment: 7 |
8/1/18, 12:00 AM | 9/25/18, 12:00 AM |
Suspect data towards the end of the deployment. Something happened in early Aug 2018 that caused the CT sensor stopped working temporarily. Issue should be investigated and annotated. A similar issue is seen in the data from CE04OSSM-SBD11-06-METBKA000, CE07SHSM-SBD11-06-METBKA000 and CE09OSSM-SBD11-06-METBKA000 from the same time period. By Lori Garzio, on 4/8/19 |
|
CE02SHSM-SBD11-06-METBKA000 Deployment: 1 |
Downwelling Shortwave Irradiance (shortwave_irradiance) and Net Shortwave Irradiance (met_netsirr) minimum values jump to ~10 W m-2 shortly after deployment (should be around 0 W m-2). A similar issue was seen on GS01SUMO-SBD11-06-METBKA000 (see annotation ID 1522 and Redmine 12543). This should be investigated and annotated. By Lori Garzio, on 4/8/19 |
||
|
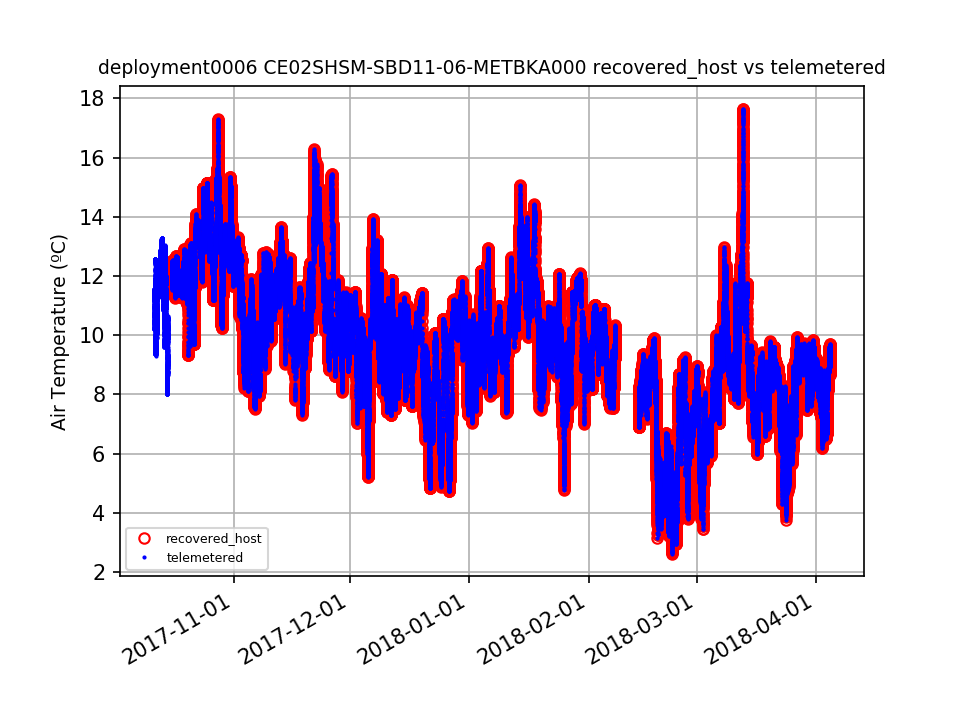
CE02SHSM-SBD11-06-METBKA000 Deployment: 6 |
2/7/18, 6:32 PM | 2/13/18, 8:35 PM |
Data gap should be annotated. By Lori Garzio, on 4/8/19 |
|
CE02SHSM-SBD11-06-METBKA000 Deployment: 6 |
10/10/17, 2:21 AM | 10/14/17, 6:34 PM | |
|
CE02SHSM-SBD11-06-METBKA000 Deployment: 5 |
4/20/17, 5:23 PM | 10/15/17, 7:06 PM |
According to annotation IDs 1430-1431, no CT data are expected. However, sea_surface_temperature values are -5, sea_surface_conductivity values are 0.0, and met_salsurf values are 0.0. It looks like other variables are using these data in their calculations, which is creating bad data products. Excluding from summary data ranges By Lori Garzio, on 4/8/19 |
|
CE02SHSM-SBD11-06-METBKA000 Deployment: 4 |
Data gaps should be annotated: [['2017-01-02T10:59:17', '2017-01-03T23:08:12'], ['2017-01-26T12:11:28', '2017-01-29T14:44:28'], ['2017-01-29T14:44:28', '2017-01-31T18:27:25'], ['2017-01-31T18:45:15', '2017-02-05T13:46:10'], ['2017-02-05T21:34:14', '2017-02-08T19:43:18'], ['2017-02-08T19:57:53', '2017-02-11T13:14:58']] By Lori Garzio, on 4/8/19 |